|
8/7/2023 0 Comments Chemdoodle 
Whilst there some advantages in having all the options visible I wonder if it might be better to have some of the options available via dropdown selections. I'm not sure I need to be able to see every bond option. The main window is the drawing board and structures can be built using either the ready made templates or point and click construction one bond at a time. By default all new atoms are carbon but the atom types can be edited using the keyboard, one nice feature is that pressing a selected letter cycles through a list of atom types, for example clicking "C" cycles through, whilst "B" cycles through. 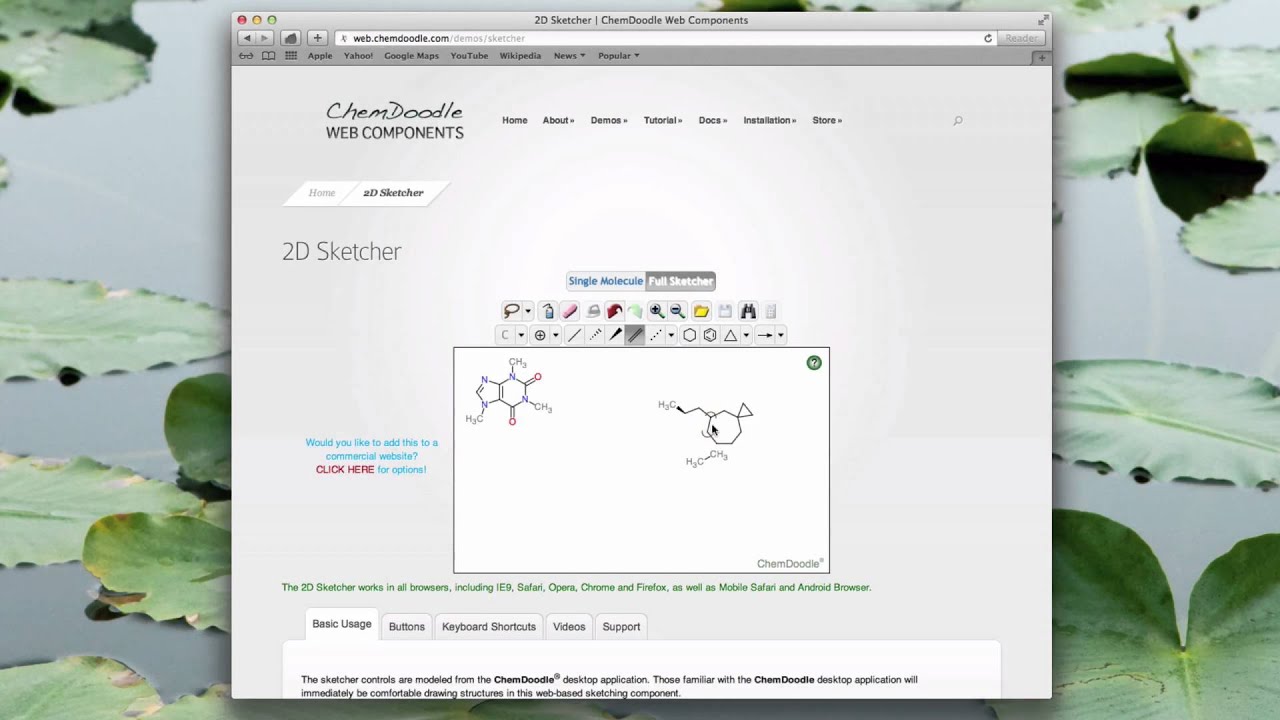
There are a number of other valuable short-cuts, e.g. Hovering over an atom and pressing "3" adds a three carbon chain. Structures can also be rapidly built using the templates available by clicking the green "Template" widget. You can add/remove hydrogens to selected atoms from the Structure menu, by default this adds hydrogens to all atoms, you can change this behaviour in the preferences but it might be useful to have an option to add hydrogens to heteroatoms only. 
Whilst the application does occupy much of the desktop one nice space saving feature is the use of tabs for each document, this means you can have multiple documents open and switch between them by simply clicking on a tab all within a single window. Whilst "Apple Z" serves as an undo function, there is also a separate history widget for each tab and you can click to a line in the history to undo/redo to the corresponding position.ĬhemDoodle provides a selection of document setting including ACS, RSC, Synthesis and Verlag Helv. All the standard chemical items such as bond types, rings, arrows, brackets are available, as well as a variety of display items such as boxes and frames. With a structure selected it is possible to generate a simulated 1H or 13C NMR spectra and paste it into the document. In fact ChemDoodle can open a variety of common chemical file types including:-ģ. 
CambridgeSoft ChemDraw Exchange (.cdx)ġ1.ISIS Sketch Transportable Graphics File (.tgf)ġ2.MDL MOLFiles, both V2000 and V3000 connection tables (.mol. sy2), Tripos Sybyl Line Notation (.sln), Beilstein ROSDAL (.ros), XYZ Files (.Mdl)ġ5.RCSB Protein Data Bank Files (.pdb. mmod), Schrödinger Maestro (.mae), Standard Molecular Data (.smd), Tripos Mol2 (.mol2. ent), RCSB Protein Data Bank Markup Language (.xml. mmcif), RCSB MacroMolecular Transmission Format (.mmtf), RCSB Protein Data Bank Files (.pdb. rd), MDL RXNFiles, both V2000 and V3000 connection tables (.rxn), MMI SketchEl Molecule (.el), Molinspiration JME String (.jme), RCSB Binar圜IF (.bcif), RCSB Macromolecular Crystallographic Information File (.cif. dx), ISIS Sketch File (.skc), ISIS Sketch Transportable Graphics File (.tgf), MDL MOLFiles, both V2000 and V3000 connection tables (.mol. smiles), IUPAC InChI (.inchi), IUPAC JCAMP-DX (.jdx. Read and write many popular chemical file types for working with the applications you use:ĪCD/ChemSketch Documents (.sk2), ChemDoodle Documents (.icl), ChemDoodle 3D Scenes (.ic3), ChemDoodle Javascript Data (.cwc.js), CambridgeSoft ChemDraw Exchange (.cdx), CambridgeSoft ChemDraw XML (.cdxml), Crystallographic Information Format (.cif), CHARMM CARD File (.crd), ChemAxon Marvin Document (.mrv), Chemical Markup Language (.cml), Daylight SMILES (.smi. Algorithmic Analysis of Cahn−Ingold−Prelog Rules of Stereochemistry: Proposals for Revised Rules and a Guide for Machine Implementation. and is 100% accurate in all 300 test cases provided. The CIP algorithm in ChemDoodle is validated against the test suite provided by Hanson et. Stereochemical features in your structures will be assigned "R", "S", "E", "Z", "M" and "P" descriptors. to remove any ambiguities and describe a completely consistent system for CIP assignments.ĬhemDoodle implements all 6 current CIP rules as well as auxilliary desciptors and mancude ring support. The most recent CIP rules from IUPAC were then algorithmically analyzed and standarized by Hanson et al. These rules were adopted by IUPAC for naming standards and fully described in the Blue books. While flawed, they have seen many revisions over the decades and were clarified by the work of Paulina Mata. The CIP rules have long been the standard for describing configurations of stereochemical features in a molecule. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
